|
Do I need to add all the identified modules, then how can I extract their names. I am not sure which modules to add in this code. Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), gene set enrichment analysis (GSEA), and weighted gene co-expression network analysis (WGCNA) were used to study the molecular mechanisms underlying diabetes and sepsis. 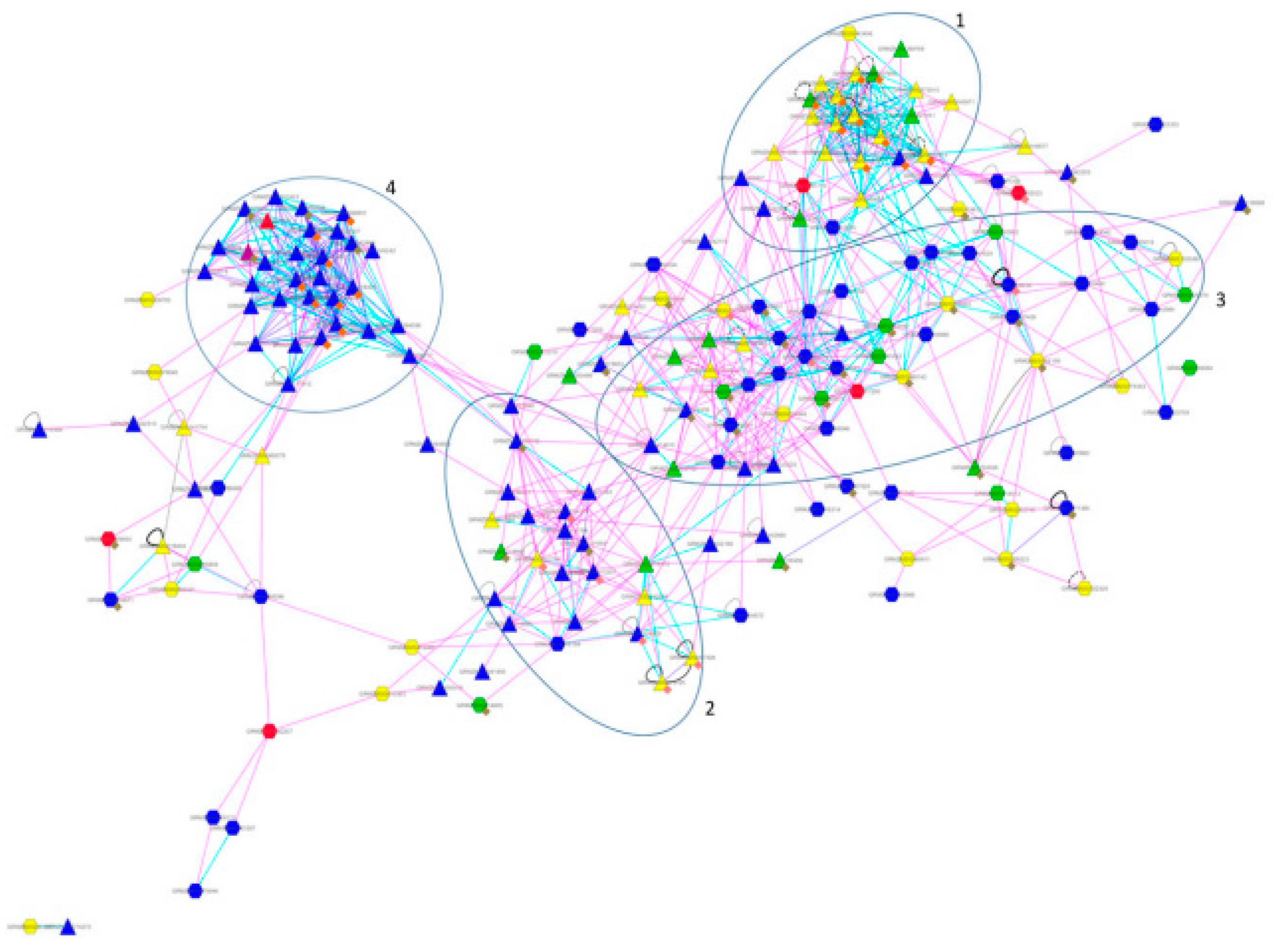
NodeFile = paste("CytoscapeInput-nodes0-", paste(modules, collapse="-"), ".txt", sep=""), InModule = is.finite(match(moduleColors, modules)) ĭimnames(modTOM) = list(modProbes, modProbes)ĮdgeFile = paste("CytoscapeInput-edges0-", paste(modules, collapse="-"), ".txt", sep=""), Load(file = "networkConstruction.RData") Īdjacency <- adjacency(datExpr, power = softPower) Export the network into edge and node list files that Cytoscape can read cyt exportNetworkToCytoscape (modadj, edgeFile './Data/cytoscape/edgessubnetwork1.txt', nodeFile './Data/cytoscape/nodessubnetwork1. I found this code to use for this purpose: load(file = "dataInput.RData") Description This function is provided by cyREST developpers and converts an igraph object into a JSON object for Cytoscape. A popular tool is Weighted Gene Correlation Network Analysis (WGCNA). Open Cytoscape and import your network by File > Import > Network from File and choose the edge-file you’ve created with Cytoscapeextract.R. I want to make a network figure using Cytoscape for all the traits. The following script will download and export files to the same directory as this.  
I have 50 traits and WGCNA analysis identified 97 modules.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
